Cellular senescence is a response to nonlethal stress that results in a persistent and often extremely stable cell cycle arrest. Among the most prominent intrinsic stimuli that induce a senescence response are genomic and epigenomic perturbations many of which can initiate or promote aging and age-related diseases. The underlying mechanisms for these perturbations are largely unknown. Senescence-inducing stimuli include progressive telomere shortening (replicative senescence) but also other stimuli that by-and-large do not impact telomere integrity such as oncogenic signaling (oncogene-induced senescence), DNA damage, oxidative stress and cytokines among others. Cultured cells usually reach senescence within several days to weeks after exposure to senescence-inducing stressors, but once arrested, they remain viable for years thereafter. To date, most studies have ignored the temporal and hierarchical evolution of senescence. However, a clear understanding of the gene regulatory modules that execute and maintain senescence will only emerge through the temporal characterization of the process. A dynamic characterization of senescence-specific gene modules and chromatin structure will reveal the nature of pathways that are activated at each stage and is thus instrumental to infer causality, to identify potential chromatin-based molecular clocks and to reveal liabilities for targeted elimination.
Addressing these challenges will be critical to develop strategies for the specific elimination of senescent cells in aging and age-related pathologies as well as for reprogramming of senescence clocks. The research I am developping in Oliver Bischof’s group aims at bridging these important knowledge gaps, charting the senescence-associated evolution and dynamics of transcription factor binding and epigenomic marks, the implications of these on gene regulation and their potential influence in patho-physiology.
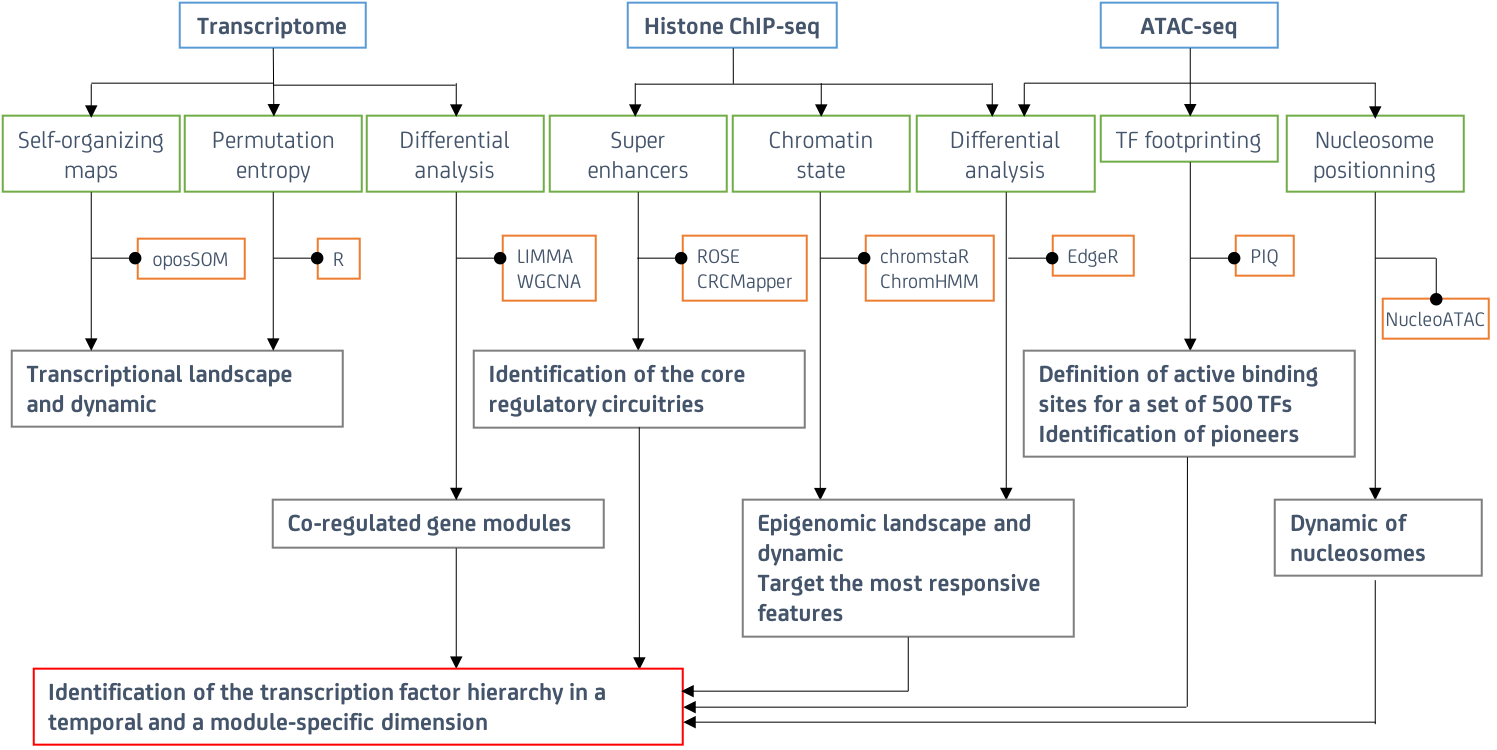
Focusing so far on oncogene-induced senescence, we probe the evolution of fibroblasts experiencing senescence using genomic approach depicting altogether the histone modification landscape (ChIP-seq for H3K4me1, H3K4me3, H3K27ac, H3K27me3), the chromatin landscape (ATAC-seq), and the transcriptionnal landscape. To deal with the the temporal and multilevel complexity of the data generated, we developped a multi-omic integrative workflow implementing cuting edge approaches for epigenomic data integration, systems biology, dimensionality reduction and variable selection (self-organizing maps, permutation entropy, chromatin state differential analysis, chromatin digital footprinting, sparse projection to latent structure discriminant analysis) summarized in the Figure 1. The information obtained can further being processed using networking approaches to understand the senescence transcriptomic dynamic in the light of epigenetic changes and transcription factor hierarchies.